Hepatovirus (taxid:12091)
VIRION
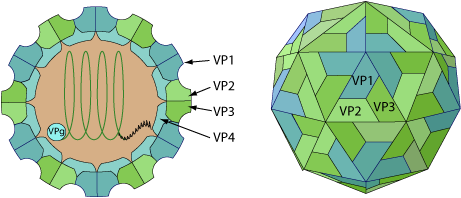
Non-enveloped, spherical, about 30 nm in diameter, T=pseudo3 icosahedral capsid surrounding the naked RNA genome. The capsid consists of a densely-packed icosahedral arrangement of 60 protomers, each consisting of 3 polypeptides, VP1, VP2, and VP3. VP4 does not seem to be incorporated into virions.
GENOME
Monopartite, linear ssRNA(+) genome of 7.478 kb, polyadenylated, composed of a single ORF encoding a polyprotein. Viral genomic RNA has a viral protein (VPg) at its 5' end instead of a methylated nucleotide cap structure. The long UTR at the 5' end contains a type III internal ribosome entry site (IRES). The P1 region encodes the structural polypeptides. The P2 and P3 regions encode the nonstructural proteins associated with replication. Encodes a unique protease 3C. The shorter 3' UTR is polyadenylated and is important in (-)strand synthesis.
GENE EXPRESSION
The virion RNA is infectious and serves as both the genome and viral messenger RNA. The IRES allows direct translation of the polyprotein. The polyprotein is initially processed by the viral protease into three precursor proteins, P1, P2, and P3. Precursor P1 is then proteolytically cleaved to yield the structural proteins. Precursors P2 and P3 are processed into replicase, VPg, and a number of proteins that modify the host cell, ultimately leading to cell lysis.
Unlike other picornaviruses, HAV needs the eIF4G factor for the initiation of translation and thus cannot shut down host protein synthesis. HAV displays highly deoptimized codon usage. The resulting slow translation rate probably contributes to the high stability of viral capsid and may play a role in escaping to host cell defenses
 .
.
ENZYMES
- RNA-dependent RNA polymerase [RdRp]
- VPG-type capping [VPg]
- NTPase-helicase [2C]
- Polyprotein major protease (Peptidase C3) [3Cpro]
REPLICATION
CYTOPLASMIC
- Attachement of the virus to host receptors mediates endocytosis of the virus into the host cell possibly by clathrin-dependent endocytosis.
- Upon endosomal acidification, the capsid undergoes a conformational change and releases VP4 that opens a pore in the host endosomal membrane and the viral genomic RNA penetrates into the host cell cytoplasm.
- VPg is removed from the viral RNA, which is then translated into a processed polyprotein.
- Replication occurs in viral factories made of membrane vesicles derived from the ER. A dsRNA genome is synthesized from the genomic ssRNA(+).
- The dsRNA genome is transcribed/replicated thereby providing viral mRNAs/new ssRNA(+) genomes.
- New genomic RNA is believed to be packaged into preassembled procapsids.
- Cell lysis and virus release.
- Maturation of provirions by an unknown host protease.
Host-virus interaction
Apoptosis modulation
Picornaviruses modulate host apoptosis  .
.
Autophagy modulation
Picornaviruses subvert the cell autophagic pathway to facilitates viral infection  :
:
Innate immune response inhibition
HAV inhibits the host IFN-mediated response by cleaving MAVS 
HAV inhibits Toll-like receptor 3 mediated antiviral response cleaving TRIF


 .
.
Matching UniProtKB/Swiss-Prot entries
(all links/actions below point to uniprot.org website)10 entries grouped by strain
1 entry
Human hepatitis A virus genotype IB (isolate HM175) (HHAV) (Human hepatitis A virus (isolate Human/Australia/HM175/1976)) reference strain
1 entry
Human hepatitis A virus genotype IA (isolate GBM) (HHAV) (Human hepatitis A virus (isolate Human/Germany/GBM/1976))
1 entry
Human hepatitis A virus genotype IA (isolate H2) (HHAV) (Human hepatitis A virus (isolate Human/China/H2/1982))
1 entry
Human hepatitis A virus genotype IA (isolate LA) (HHAV) (Human hepatitis A virus (isolate Human/Northern California/LA/1974))
1 entry
Human hepatitis A virus genotype IB (isolate MBB) (HHAV) (Human hepatitis A virus (isolate Human/Northern Africa/MBB/1978))
1 entry
Human hepatitis A virus genotype IIA (isolate CF-53) (HHAV) (Human hepatitis A virus (isolate Human/France/CF-53/1979))
1 entry
Human hepatitis A virus genotype IIB (isolate SLF88) (HHAV) (Human hepatitis A virus (isolate Human/Sierra Leone/SLF88/1988))
1 entry
Human hepatitis A virus genotype IIIA (isolate NOR-21) (HHAV) (Human hepatitis A virus (isolate Human/Norway/NOR-21/1998))
1 entry
Human hepatitis A virus genotype IIIB (isolate HAJ85-1) (HHAV) (Human hepatitis A virus (isolate Human/Japan/HAJ85-1/1985))
1 entry
Simian hepatitis A virus genotype V (isolate AGM-27) (SHAV) (Simian hepatitis A virus (isolate Cercopithecus/Kenya/AGM-27/1985))
hepatovirus B1 taxid:1708572
| Protein | ModelArchive |
| Genome polyprotein | ma-jd-viral-30863 |
