Restriction-modification system evasion by virus (kw:KW-1258)
The restriction-modification (RM) system is a defense mechanism present in over 90% of sequenced bacterial and archeal genomes. Its consists of a modification enzyme that methylates a specific DNA sequence in a genome and a restriction endonuclease that cleaves DNA lacking this methylation  .<div id="PMID: 11782494" class="hidden">2001 Fred Griffith review lecture. Immigration control of DNA in bacteria: self versus non-self
.<div id="PMID: 11782494" class="hidden">2001 Fred Griffith review lecture. Immigration control of DNA in bacteria: self versus non-self
Noreen E Murray
Microbiology (Reading, Engl.) January 2002; 148: 3?20
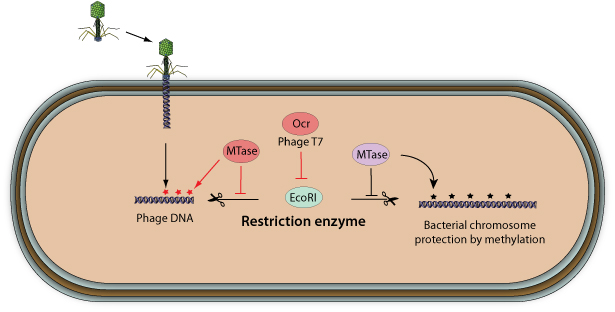
Bacterial viruses have evolved different strategies to evade the restriction-modification system 
 . Some viruses encode their own methyltransferase in order to protect their genome from host restriction enzymes
. Some viruses encode their own methyltransferase in order to protect their genome from host restriction enzymes  . Instead, bacteriophage T7 encodes the OCR protein that blocks the active site of several restriction enzymes by mimicking the phosphate backbone of B-form DNA. Other bacterial viruses use unusual bases on their genome to avoid restriction. Bacteriophages SPO1, SP82, and 2C replace thymidine with 5-hydroxymethyluracil while phages PBS1 and PBS2 thymine is completely changed to uracil.
. Instead, bacteriophage T7 encodes the OCR protein that blocks the active site of several restriction enzymes by mimicking the phosphate backbone of B-form DNA. Other bacterial viruses use unusual bases on their genome to avoid restriction. Bacteriophages SPO1, SP82, and 2C replace thymidine with 5-hydroxymethyluracil while phages PBS1 and PBS2 thymine is completely changed to uracil.
| Family | Genus | Virus | Strategy of RM evasion | Viral protein | Ref. |
| Siphoviridae | Tunalikevirus | Bacteriophage T1 | Viral methyltransferase | Dmt methyltransferase |   |
| Podoviridae | T7likevirus | Bacteriophage T3 | Viral SAMase | S-adenosyl-L-methionine hydrolase |  |
| Myoviridae | T4likevirus | Bacteriophage T4 | Viral methyltransferase | Dam methyltransferase |  |
| Myoviridae | T4likevirus | Bacteriophage T2 | Viral methyltransferase | Dam methyltransferase |  |
| Podoviridae | T7likevirus | Bacteriophage T7 | Inhibition of type-I RM enzymes | OCR protein |  |
| Myoviridae | Punalikevirus | Bacteriophage P1 | Internal capsid protein | DarA, DarB |  |
| Podoviridae | P22likevirus | Bacteriophage P22 | Modification of host methylase activity | Ral |  |
| Siphoviridae | Lambdalikevirus | Bacteriophage λ reverse | Modification of host methylase activity | Lar |  |
| Siphoviridae | Lambdalikevirus | Bacteriophage λ | Modification of host methylase activity | Ral |  |
| Myoviridae | Mulikevirus | Bacteriophage Mu | Adenine acetyltransferase | Mom |   |
Michael R Hall, Edward McGillicuddy, Lewis J Kaplan
Surg Infect (Larchmt) January 29, 2014;
Simon J Labrie, Julie E Samson, Sylvain Moineau
Nat. Rev. Microbiol. May 2010; 8: 317-327
Julie E Samson, Alfonso H Magadan, Mourad Sabri, Sylvain Moineau
Nat. Rev. Microbiol. October 2013; 11: 675-687
B Auer, M Schweiger
J. Virol. February 1984; 49: 588-590
F W Studier, N R Movva
J. Virol. July 1976; 19: 136-145
V G Kossykh, S L Schlagman, S Hattman
J. Biol. Chem. June 16, 1995; 270: 14389-14393
C Atanasiu, T-J Su, S S Sturrock, D T F Dryden
Nucleic Acids Res. September 15, 2002; 30: 3936-3944
S Iida, M B Streiff, T A Bickle, W Arber
Virology March 1987; 157: 156-166
A V Semerjian, D C Malloy, A R Poteete
J. Mol. Biol. May 5, 1989; 207: 1-13
G King, N E Murray
Gene May 19, 1995; 157:225
I TAKAHASHI, J MARMUR
Nature February 23, 1963; 197: 794-795
Swinton D, Hattman S, Crain PF, Cheng CS, Smith DL, McCloskey JA
Proc Natl Acad Sci U S A. 1983 Dec;80(24):7400-4
James Murphy, Jennifer Mahony, Stuart Ainsworth, Arjen Nauta, Douwe van Sinderen
Appl Environ Microbiol December 2013; 79: 7547-7555
E. F. Wagner, B. Auer, M. Schweiger
J. Virol. March 1979; 29: 1229-1231
V. G. Kossykh, S. L. Schlagman, S. Hattman
J. Bacteriol. May 1997; 179: 3239-3243
Naofumi Handa, Ichizo Kobayashi
J. Bacteriol. November 2005; 187: 7362-7373
W. A. Loenen, N. E. Murray
J. Mol. Biol. July 5, 1986; 190: 11-22
A. Toussaint
Virology March 1976; 70: 17-27
F. W. Studier
J. Mol. Biol. May 15, 1975; 94: 283-295
Matching UniProtKB/Swiss-Prot entries
(all links/actions below point to uniprot.org website)45 entries grouped by strain
7 entries
Enterobacteria phage T4 (Bacteriophage T4) reference strain
4 entries
Pseudomonas phage PaMx11 reference strain
4 entries
Salmonella phage ViI reference strain
3 entries
Delftia phage PhiW-14 (Deftia acidovorans bacteriophage phiW-14) reference strain
3 entries
Escherichia phage P1 (Bacteriophage P1) reference strain
1 entry
Bacillus phage SP01 (Bacteriophage SP01) reference strain
1 entry
Bacillus phage SPbeta (Bacillus phage SPBc2) (Bacteriophage SP-beta) reference strain
1 entry
Escherichia phage Mu (Bacteriophage Mu) reference strain
1 entry
Escherichia phage RB69 (Bacteriophage RB69) reference strain
1 entry
Escherichia phage T1 (Bacteriophage T1) reference strain
1 entry
Escherichia phage T7 (Bacteriophage T7) reference strain
1 entry
Escherichia phage lambda (Bacteriophage lambda) reference strain
1 entry
Haemophilus phage HP1 (strain HP1c1) (Bacteriophage HP1) reference strain
1 entry
Salmonella phage P22 (Bacteriophage P22) reference strain
6 entries
Pseudomonas phage M6
2 entries
Enterobacteria phage T2 (Bacteriophage T2)
1 entry
Bacillus phage SP10 (Bacillus phage SP-10)
1 entry
Bacillus phage SPR (Bacteriophage SPR)
1 entry
Bacillus phage phi3T (Bacteriophage phi-3T)
1 entry
Bacillus phage rho11s (Bacteriophage rho-11s)
1 entry
Enterobacteria phage T6 (Bacteriophage T6)
1 entry
Enterobacteria phage phi21 (Bacteriophage phi-21)
1 entry
