Viral receptor tropism switching (kw:KW-1264)
Some bacteriophages have developed tropism-switching genetic cassettes, that allow them to modify their fibers proteins, and thus to bind to a broader range of hosts.
Different mechanisms can be used:
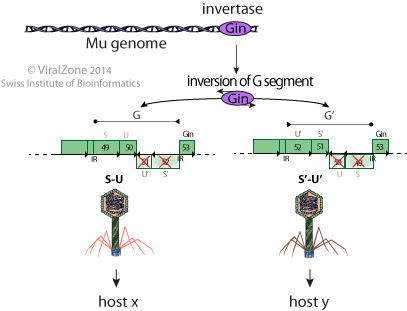
- The host range variation system found in phages such as Mu and P1 is a genetic system that allows some bacterial viruses to switch between two types of fibers by inversion of the DNA segment encoding the fibers. The change in fibers specificity is performed through site specific inversion of a genome segment catalyzed by a virally encoded invertase. This genome segment codes for two alternate sets of tail fibers, of which only the "well oriented" one is expressed
 .
. - Bordetella bacteriophages generate diversity in a gene that specifies host tropism produced through a genetic element that combines the basic retroelement
life cycle of transcription, reverse transcription and integration with site-directed, adenine-specific mutagenesis
 .
.
Tropism switching in Bordetella bacteriophage defines a family of diversity-generating retroelements
Sergei Doulatov, Asher Hodes, Lixin Dai, Neeraj Mandhana, Minghsun Liu, Rajendar Deora, Robert W. Simons, Steven Zimmerly, Jeff F. Miller
Nature September 23, 2004; 431: 476-481
Sergei Doulatov, Asher Hodes, Lixin Dai, Neeraj Mandhana, Minghsun Liu, Rajendar Deora, Robert W. Simons, Steven Zimmerly, Jeff F. Miller
Nature September 23, 2004; 431: 476-481
A genetic switch in vitro: DNA inversion by Gin protein of phage Mu
R. H. Plasterk, R. Kanaar, P. van de Putte
Proc. Natl. Acad. Sci. U.S.A. May 1984; 81: 2689-2692
R. H. Plasterk, R. Kanaar, P. van de Putte
Proc. Natl. Acad. Sci. U.S.A. May 1984; 81: 2689-2692
Matching UniProtKB/Swiss-Prot entries
(all links/actions below point to uniprot.org website)10 entries grouped by strain
5 entries
Escherichia phage Mu (Bacteriophage Mu) reference strain
3 entries
Escherichia phage P1 (Bacteriophage P1) reference strain
1 entry
Bordetella phage BPP-1 reference strain
1 entry
