Lymphocryptovirus (taxid:10375)
VIRION
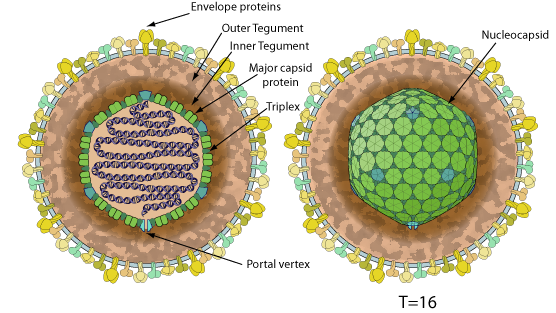
Enveloped, spherical to pleomorphic, 150-200 nm in diameter, T=16 icosahedral symmetry. Capsid consists of 162 capsomers and is surrounded by an amorphous tegument. Glycoproteins complexes are embedded in the lipid envelope.
GENOME
Monopartite, linear, dsDNA genome of about 180 kb. The genome contains terminal and internal reiterated sequences.
GENE EXPRESSION
Each viral transcript usually encodes a single protein and has a promoter/regulatory sequence, a TATA box, a transcription initiation site, a 5' leader sequence of 30-300 bp (not translated), a 3' untranslated sequence of 10-30 bp, and a poly A signal. There are many gene overlaps. There are only few spliced genes. Some of the expressed ORFs are antisense to each other. Some ORFs can be accessed from more than one promoter. There are some non-coding genes.
ENZYMES
- DNA-dependent DNA polymerase
- DNA primase
- Tegument deneddylase (Peptidase C76)
- Assemblin (Peptidase S21)
- Kinase
- Helicase
- Ribonucleoside-diphosphate reductase
- Thymidine kinase
- Uracil-DNA glycosylase
REPLICATION
NUCLEAR
Lytic replication:
- Attachment of the viral glycoproteins to host receptors mediates endocytosis of the virus into the host cell.
- Fusion with the plasma membrane to release the core and the tegument proteins into the host cytoplasm.
- The capsid is transported to the nuclear pore where the viral DNA is released into the nucleus.
- Transcription of immediate early genes which promote transcription of early genes and protect the virus against innate host immunity.
- Transcription of early viral mRNA by host polymerase II, encoding proteins involved in replication of the viral DNA.
- A first round of circular genome amplification occurs by bidirectional replication
- Synthesis of linear concatemer copies of viral DNA by rolling circle.
- Transcription of late mRNAs by host polymerase II, encoding structural proteins.
- Assembly of the virus in nuclear viral factories and budding through the inner lamella of the nuclear membrane which has been modified by the insertion of herpes glycoproteins, throughout the Golgi and final release at the plasma membrane.
Latent replication : replication of circular viral episome in tandem with the host cell DNA using the host cell replication machinery.
Host-virus interaction
Adaptive immune response inhibition
Epstein-Barr virus protein BNLF2a inhibits host adaptive immune response by interacting with TAP1 and TAP2 and thus preventing TAP-mediated peptide transport and subsequent loading. The glycine-alanine repeat domain (GAr) of Epstein-Barr virus-encoded nuclear antigen 1 (EBNA1) prevents major histocompatibility complex (MHC) class I-restricted presentation of EBNA1 epitopes to cytotoxic T cells.
Apoptosis modulation
Epstein-Barr viral protein BHRF1 is a viral homologue of the Bcl-2 oncogene that is able to protect cells from induced apoptosis. EBNA3C directly interacts with host Gemin3 and thereby blocks p53-mediated apoptosis  .
.
Autophagy modulation
The viral Bcl-2 homolog BHRF1 in addition to prevent host cel death, may also control host autophagy  .
.
Cell-cycle modulation
EBNA3C forms a complex with Cyclin D1 thereby facilitating the G1-S phase transition  . Additionally, EBNA2 can affect activities of cell cycle regulators and retards cell cycle progression at G2/M phase.
The conserved UL24 family of human alpha, beta and gamma herpesviruses induces a cell cycle arrest at G2/M transition through inactivation of the host cyclinB/cdc2 complex. EBV encodes an UL24 homolog termed BXRF1 that should perform this role
. Additionally, EBNA2 can affect activities of cell cycle regulators and retards cell cycle progression at G2/M phase.
The conserved UL24 family of human alpha, beta and gamma herpesviruses induces a cell cycle arrest at G2/M transition through inactivation of the host cyclinB/cdc2 complex. EBV encodes an UL24 homolog termed BXRF1 that should perform this role  .
.
Innate immune response inhibition
EBV inhibits the cascade leading to production of interferon-beta by targeting host IRF3 protein with the BGLF4 protein kinase that directly interacts with and inhibits host IRF3. BRLF1 expression also decreases induction of IFN-beta, and reduces expression of IRF3 and IRF7.
Host splicing inhibition
EBV EB2 modulates the host mRNA expression by exporting unspliced mRNA, thereby inducing alternative splicing  .
.
Matching UniProtKB/Swiss-Prot entries
(all links/actions below point to uniprot.org website)196 entries grouped by protein
3 entries
Shutoff alkaline exonuclease (SOX) (EC 3.1.-.-)
2 entries
Apoptosis regulator BALF1
3 entries
Secreted protein BARF1 (33 kDa early protein) (p33)
1 entry
Late gene expression regulator BDLF3.5
3 entries
Protein BDLF2
1 entry
Glycoprotein BDLF3
1 entry
Late gene expression regulator BFRF2
1 entry
Uncharacterized protein BHLF1
1 entry
G-protein coupled receptor BILF1 (GPCR BILF1)
1 entry
Glycoprotein BILF2
3 entries
Tegument protein BKRF4
3 entries
Uncharacterized protein BLLF2
3 entries
Tegument protein BLRF2
3 entries
Protein BMRF2
3 entries
Protein BNLF2a
3 entries
Uncharacterized protein BNLF2b
1 entry
Replication and transcription activator (Rta) (Immediate-early protein Rta) (Protein R)
2 entries
Replication and transcription activator (Rta) (Immediate-early protein Rta) (Protein R)
3 entries
Transcriptional activator BRRF1
3 entries
Tegument protein BRRF2
3 entries
Uncharacterized protein BTRF1
1 entry
Late gene expression regulator BVLF1
3 entries
Lytic switch protein BZLF1 (EB1) (Protein Z) (Trans-activator protein BZLF1) (Zebra) (Zta) (bZIP transcription factor ZEBRA)
3 entries
Cytoplasmic envelopment protein 1
3 entries
Cytoplasmic envelopment protein 2 (Tegument protein BGLF2)
2 entries
Cytoplasmic envelopment protein 3
1 entry
Capsid vertex component 1
3 entries
Capsid vertex component 2
3 entries
Major DNA-binding protein
3 entries
DNA polymerase catalytic subunit (EC 2.7.7.7)
3 entries
Deoxyuridine 5'-triphosphate nucleotidohydrolase (dUTPase) (EC 3.6.1.23) (dUTP pyrophosphatase)
3 entries
DNA polymerase processivity factor BMRF1 (Early Antigen Diffused) (EA-D) (Polymerase accessory subunit)
3 entries
Apoptosis regulator BHRF1 (Early antigen protein R) (EA-R) (Nuclear antigen)
3 entries
Epstein-Barr nuclear antigen 1 (EBNA-1) (EBV nuclear antigen 1) (EC 3.1.21.-)
3 entries
Epstein-Barr nuclear antigen 2 (EBNA-2) (EBV nuclear antigen 2)
3 entries
Epstein-Barr nuclear antigen 3 (EBNA-3) (EBV nuclear antigen 3) (Epstein-Barr nuclear antigen 3A) (EBNA-3A) (EBV nuclear antigen 3A)
3 entries
Epstein-Barr nuclear antigen 4 (EBNA-4) (EBV nuclear antigen 4) (Epstein-Barr nuclear antigen 3B) (EBNA-3B) (EBV nuclear antigen 3B)
3 entries
Epstein-Barr nuclear antigen leader protein (EBNA-LP) (EBV nuclear antigen leader protein) (Epstein-Barr nuclear antigen 5) (EBNA-5) (EBV nuclear antigen 5)
3 entries
Epstein-Barr nuclear antigen 6 (EBNA-6) (EBV nuclear antigen 6) (Epstein-Barr nuclear antigen 3C) (EBNA-3C) (EBV nuclear antigen 3C) (Epstein-Barr nuclear antigen 4B) (EBNA-4B) (EBV nuclear antigen 4B)
3 entries
Envelope glycoprotein B (gB)
3 entries
Envelope glycoprotein H (gH)
3 entries
Envelope glycoprotein L (gL)
3 entries
Envelope glycoprotein M (gM)
3 entries
Envelope glycoprotein N
3 entries
Envelope glycoprotein GP350 (Membrane antigen) (MA)
3 entries
Glycoprotein 42 (gp42)
1 entry
DNA replication helicase (EC 3.6.4.-)
1 entry
DNA helicase/primase complex-associated protein (HEPA) (Primase-associated factor)
3 entries
mRNA export factor ICP27 homolog (Mta) (ORF57 protein homolog) (Protein EB2) (Protein SM)
3 entries
Viral interleukin-10 homolog (vIL-10) (20 kDa protein) (Protein BCRF1)
3 entries
Inner tegument protein
3 entries
Thymidine kinase (EC 2.7.1.21)
3 entries
Serine/threonine-protein kinase BGLF4 (EC 2.7.11.1)
3 entries
Uncharacterized LF1 protein
3 entries
Protein LF2
2 entries
Uncharacterized protein LF3
5 entries
Latent membrane protein 1 (LMP-1) (Protein p63)
3 entries
Latent membrane protein 2 (Terminal protein)
3 entries
Large tegument protein deneddylase (EC 3.4.19.12) (EC 3.4.22.-)
3 entries
Major capsid protein (MCP)
3 entries
Major tegument protein (MTP) (Protein p140)
2 entries
Nuclear egress protein 1
1 entry
Nuclear egress protein 2
3 entries
Portal protein
1 entry
DNA primase (EC 2.7.7.-)
3 entries
Ribonucleoside-diphosphate reductase large subunit (R1) (EC 1.17.4.1) (Ribonucleotide reductase large subunit)
3 entries
Ribonucleoside-diphosphate reductase small subunit (EC 1.17.4.1) (Ribonucleotide reductase small subunit)
2 entries
Uncharacterized protein RPMS1
3 entries
Capsid scaffolding protein (Capsid protein P40) (Protease precursor) (pPR) (Protein EC-RF3/EC-RF3A) (Virion structural protein BVRF2)
1 entry
Small capsomere-interacting protein
3 entries
Tegument protein UL51 homolog
1 entry
Tripartite terminase subunit 1
3 entries
Tripartite terminase subunit 3 (EC 3.1.-.-) (Terminase large subunit)
1 entry
Triplex capsid protein 1
1 entry
Triplex capsid protein 2
1 entry
Protein UL24 homolog
1 entry
Packaging protein UL32 homolog
1 entry
Late gene expression regulator BcRF1 (TBP-like protein BcRF1)
2 entries
Late gene expression regulator BDLF4
1 entry
Late gene expression regulator BGLF3
1 entry
Uracil-DNA glycosylase (UDG) (EC 3.2.2.27) (UNG)
1 entry
Envelope glycoprotein GP340 (Membrane antigen) (MA)
1 entry
Uncharacterized protein EC-RF4
Epstein-Barr virus taxid:10376
Epstein-Barr virus (strain AG876) taxid:82830
Epstein-Barr virus (strain B95-8) taxid:10377
Human herpesvirus 4 type 2 taxid:12509
| Protein | ModelArchive |
| Latent membrane protein 1 (Protein p63) | ma-jd-viral-30641 |
