Merhavirus (taxid:2842863)
VIRION
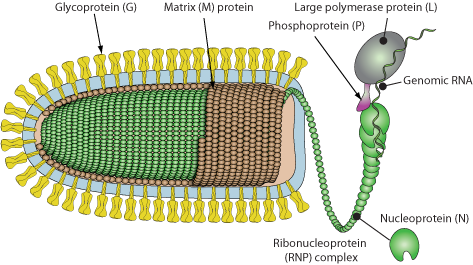
Unknown, discovered using viral metagenomics. Presumably enveloped, bullet shaped.
GENOME
Negative-stranded RNA linear genome, about 11.5kb in size. Encodes for five proteins. Enzymes: RNA dependent RNA polymerase, RNA guanylyl transferase, mRNA methyltransferase.
GENE EXPRESSION
The viral RNA dependent RNA polymerase binds the encapsidated genome at the leader region, then sequentially transcribes each genes by recognizing start and stop signals flanking viral genes. mRNAs are capped and polyadenylated by the L protein during synthesis.
ENZYMES
- RNA-dependent RNA polymerase L
- Mononega-type capping
- RNA TPase, GTase, N7 Mtase, 2'O Mtase L
REPLICATION
CYTOPLASMIC
- Attachment of the viral G glycoproteins to host receptors mediates endocytosis of the virus into the host cell.
- Fusion of virus membrane with the vesicle membrane; ribonucleocapsid is released into the cytoplasm.
- Sequential transcription , viral mRNAs are capped and polyadenylated by polymerase stuttering in the cytoplasm.
- Replication presumably starts when enough nucleoprotein is present to encapsidate neo-synthetized antigenomes and genomes.
- The ribonucleocapsid binds to the matrix protein and buds, releasing new virions.
Culex tritaeniorhynchus rhabdovirus taxid:936308
| Protein | ModelArchive |
| Glycoprotein | ma-jd-viral-07054 |
| Matrix protein | ma-jd-viral-03356 |
| Nucleoprotein (Nucleocapsid protein) | ma-jd-viral-15095 |
| Phosphoprotein | ma-jd-viral-40129 |
