Protoparvovirus (taxid:1506574)
VIRION
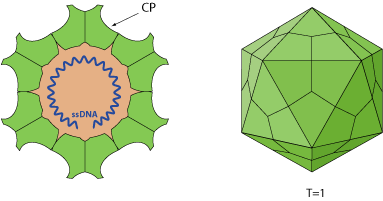
Non-enveloped, round, T=1 icosahedral symmetry, 18-26 nm in diameter. The capsid consists of 60 copies of CP protein.
GENOME
Linear, ssDNA genome of about 4 to 6 kb in size. Both positive and negative strands are encapsidated, although the percentage of particles encapsidating the positive strand can be lower depending on the host cell. MVM packages predominantly negative-sense single strands, while LuIII encapsidates strands of both polarities with equal efficiency. ORFs for both the structural and non-structural proteins are located on the same DNA strand.
The genome is replicated through rolling-hairpin mechanism.
GENE EXPRESSION
Host proteins transcribe the genomes into mRNAs. Alternative splicing allows expression of both structural and non-structural proteins.
ENZYMES
REPLICATION
NUCLEAR
- Attachement to host receptors initiates clathrin-mediated endocytosis of the virion into the host cell.
- The virion penetrates into the cytoplasm via permeabilization of host endosomal membrane.
- Microtubular transport of the virion toward the nucleus.
- The viral ssDNA genome penetrates into the nucleus.
- The ssDNA is converted into dsDNA by cellular proteins.
- dsDNA transcription gives rise to viral mRNAs when host cell enters S phase and translated to produce viral proteins.
- Replication occurs through rolling-hairpin mechanism, with NS1 endonuclease binding covalently to the 5' genomic end.
- Individual ssDNA genomes are excised from replication concatemers by a process called junction resolution.
- These newly synthesized ssDNA can either
a) be converted to dsDNA and serve as a template for transcription/replication
b) be encapsidated to form new virions that are released by cell lysis.
Host-virus interaction
Apoptosis modulation
Parvovirus infection induces host cell death in cancerous cells. Apoptosis is the major form of cell death. However, necrosis has also been reported during infection of the minute virus of mice and parvovirus H-1
 .
.
H-1PV NS1 protein expression induces apoptosis via caspases activation and DNA damage signaling. Depending on cell type and growth conditions, H-1PV is able to induce apoptosis, necrosis, or cathepsin B-dependent cell death.


Cell-cycle modulation
H-1PV NS1 protein induces G2/M checkpoint arrest.


Matching UniProtKB/Swiss-Prot entries
(all links/actions below point to uniprot.org website)25 entries grouped by strain
3 entries
Murine minute virus (strain MVM prototype) (MVM) (Murine minute virus (strain MVM(p))) reference strain
2 entries
Canine parvovirus type 2 (isolate Dog/United States/CPV-N/1978) (CPV-2) reference strain
2 entries
Feline panleukopenia virus (strain 193) (FPV)
2 entries
Hamster parvovirus H1
2 entries
Mink enteritis virus (strain Abashiri) (MEV)
2 entries
Murine minute virus (strain MVMi) (MVM) (Murine parvovirus)
2 entries
Parvovirus LuIII
2 entries
Porcine parvovirus (strain Kresse) (PPV)
2 entries
Porcine parvovirus (strain NADL-2) (PPV)
1 entry
Canine parvovirus type 2 (CPV-2)
1 entry
Canine parvovirus type 2 (isolate Dog/United States/CPV-b/1978) (CPV-2)
1 entry
Canine parvovirus type 2 (strain Dog/United States/780929/-) (CPV-2)
1 entry
Canine parvovirustype 2 (isolate Dog/United States/CPV-d/1988) (CPV-2)
1 entry
Feline panleukopenia virus (FPV)
1 entry
Porcine parvovirus (strain 90HS) (PPV)
Megabat bufavirus 1 taxid:1756191
| Protein | ModelArchive |
| Capsid protein VP1 (Coat protein VP1) | ma-jd-viral-07369 |
| Capsid protein VP2 | ma-jd-viral-07372 |
| Initiator protein NS1 (EC 3.6.4.12) (Non-structural protein NS1) | ma-jd-viral-23456 |
Porcine parvovirus 5 taxid:1241957
| Protein | ModelArchive |
| Capsid protein VP1 (Coat protein VP1) | ma-jd-viral-41339 |
| Replicase | ma-jd-viral-23500 |
Rat bufavirus SY-2015 taxid:1763507
| Protein | ModelArchive |
| Capsid protein VP1 (Coat protein VP1) | ma-jd-viral-07357 |
| Initiator protein NS1 (EC 3.6.4.12) (Non-structural protein NS1) | ma-jd-viral-23462 |
| VP2 | ma-jd-viral-07370 |
Sea otter parvovirus 1 taxid:1882382
| Protein | ModelArchive |
| Initiator protein NS1 (EC 3.6.4.12) (Non-structural protein NS1) | ma-jd-viral-23461 |
| VP1 | ma-jd-viral-07373 |
| VP2 | ma-jd-viral-07376 |
